Engineering Biology & Materials Science
Synthesis
Synthesis considers the primary production or creation of material components via engineered biology.
Introduction
We consider Synthesis as the primary production or creation of material components. In this roadmap, the focus is on the generation of material components via engineered biology. This includes utilizing or exploiting engineered biology to produce monomers, polymers, biomolecules, and macromolecules that serve as components of a material (bulk or otherwise). Key challenges in Synthesis include high-yield production of protein-based natural materials, such as spider silk and elastin, from engineered biology; alternative biological production systems and utilization of natural and non-natural nucleic and amino acids; and enabling re-synthesis or recycling of materials to enable greater sustainability.
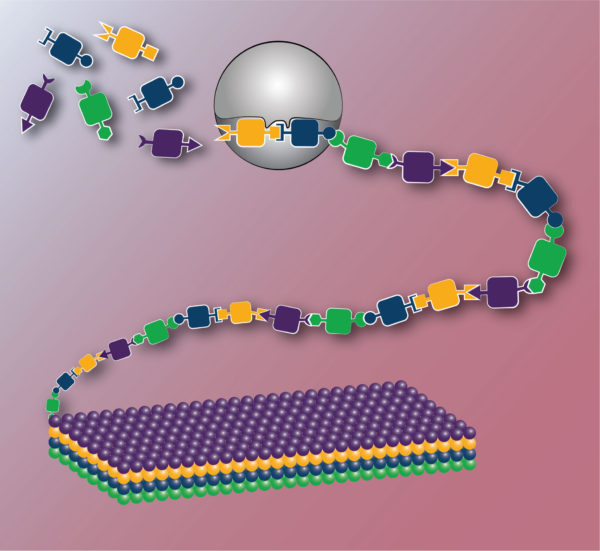
Engineering biology has the capability to transform the materials generated through biosynthesis and the catalysts which facilitate their formation. In this depiction, novel monomers with orthogonal reactivity enable sequence-defined polymerization and the precise control of monomer order provides programmed higher order structure. Equally important, this synthesis is carried out by robustly engineered enzymes able to recognize non-natural monomers.
Breakthrough Capabilities
Biological synthesis and/or polymerization of non-natural and/or abiotic chemical monomers, excluding amino acids., ,
Identify and engineer enzymes to recognize and polymerize conventional chemical monomers (e.g., acrylamide).
Bottleneck/Challenge: Limited library of employable enzymes that can be used to produce polymers in a variety of environments.
Potential Solution: Identify the target, based on protein structure and known biological roles, and develop screening strategy for directed-evolution.
Develop metabolic pathways to yield chiral cyclic monomers amenable to ring-opening polymerization.
Bottleneck/Challenge: Compatibility of aromatic, cyclic monomers with cell interior may require cell-free synthetic approaches to be developed.
Potential Solution: Engineering of synthetic lipid bilayer systems for synthesis processes.
Bottleneck/Challenge: Prediction of materials properties lags behind protein folding, inorganic material property forecasting.
Potential Solution: Requires continued investment in computational models of soft materials, meso-scale tool development for capturing impacts of processing conditions and kinetically trapped structures.
Demonstrate side chain modification of polymeric scaffolds by biological reactions catalyzed by living cells.
Bottleneck/Challenge: Dissolving conventional polymers in a condition where an enzyme can operate (i.e., temperature, oxygen, buffer, cofactors like ATP); in order for this to work, the polymer chains should dissolve well in a medium (ideally water) so that enzymes can approach.
Potential Solution: De novo engineering of an enzyme that is functional in multiple environments, such as design of an active site that doesn’t require extensive folding structure in water.
Potential Solution: “Harden” enzymes to maintain activity in non-aqueous environments.
Bottleneck/Challenge: Development of strain chassis that can access and modify polymeric scaffolds under survivable conditions.
Potential Solution: Engineering of extracellular matrix and enzyme secretion pathways to enable modification of polymeric scaffolds outside of cells.
Enable vinyl monomer production by engineering metabolic pathways for in situ polymerization reactions.
Bottleneck/Challenge: High-yield production of vinyl monomer by enzymatic cascade has yet to be achieved.
Potential Solution: Targeted directed evolution of enzymes.
Expand the library of chemical monomers polymerizable by evolved enzymes.
Bottleneck/Challenge: Testing enzyme variants in vivo is slow due to transformation and cell growth requirements.
Potential Solution: Cell-free prototyping of enzyme variants allows much higher-throughput exploration of protein space for identification of variants with favorable functionalities.
Engineer complex orthogonal translation systems, including engineered orthogonal ribosomes, for site-directed incorporation of novel (non-amino acid) chemistries into polymers.
Bottleneck/Challenge: Ribosomes do not natively polymerize most non L-α-amino-acid substrates.
Potential Solution: Use RNA modeling/design to remodel the peptidyl transferase center (PTC) of the ribosome to accommodate new amino acids.
Potential Solution: Evolve the ribosome toward incorporation of new chemistries in a cell-free environment.
Bottleneck/Challenge: Requires construction of well-coordinated biological systems in cells to function effectively.
Potential Solution: Development of genome-engineering tools that improve the throughput of designing, building, and testing effective orthogonal translation systems.
Potential Solution: Use and develop new genomic engineering and molecular evolution strategies to permit continuous diversification of biomolecular systems and multi-gene pathways for systems engineering and optimization.
Enable complex polymer production (e.g., co-polymers, sequence-controlled polymers, brushed polymers) with enzymatic systems.
Bottleneck/Challenge: Monomer and polymer solubility limit potential for bio-catalysis, especially in aqueous systems.
Potential Solution: Explore cell-free or bi-phasic conditions to drive reactions in organic solvents or at the water-solvent interface.
Bottleneck/Challenge: Inability to produce sequence-controlled polymers with both natural and synthetic chemistries.
Potential Solution: Engineer biological systems that can polymerize materials with biological and abiological monomers, such as organisms with open coding channels, engineered translation systems that can encode diverse monomers, engineered ribosomes that can catalyze new bond formation and polymerization, and synthetic organelles that can serve as dedicated sites for polymerization of synthetic biomaterials isolated from the cell and prevent cytotoxicity.
Hybrid chemical and biological synthesis methods.,
Characterize and improve the enzymology of orthogonal translation systems (OTSs) for site-directed incorporation of novel chemistries into polymers.
Bottleneck/Challenge: A variety of OTSs exist for the incorporation of over 100 non-standard chemistries into proteins; however, their utility for applications remains unclear.
Potential Solution: Perform thorough characterization of OTS expression, solubility, enzymology, for Km, Kcat, and orthogonality.
Potential Solution: Engineer enhanced orthogonality, specificity, and efficiency into OTS to permit multi-site incorporation of two or more synthetic chemistries into the same polymer.
Enable hybrid flow chemistry-biocatalytic synthesis systems.
Bottleneck/Challenge: Limited library of catalytic enzymes that would be stable at high temperatures and/or in the absence of water.
Potential Solution: Design thermophilic catalytic enzymes for use in flow reactors.
Potential Solution: Explore alternative solvents as stabilization agents (e.g., ionic liquids).
Full integration of heterogeneous chemical and biological catalysis in modular platforms that are stable after long-term storage and facilitate product switching.
Bottleneck/Challenge: Long-term storage of biological entities typically requires a cold chain for stability, thus more robust systems that realize long-term storage in ambient temperature and humidity range is needed.
Potential Solution: Innovations in genetic engineering of cell-based and cell-free transcription/translation, consortia and hybrid biological platforms to enable manufacturing flexibility.
Enable retrobiosynthesis of materials with desired properties, such as thermal conductivity or elasticity.
Bottleneck/Challenge: Mapping of traditional materials properties to biomolecules and self-assembled/hybrid systems.
Potential Solution: Databases with materials properties for characterized systems.
Bottleneck/Challenge: Paucity of data about emergent properties of materials from biocomponents.
Potential Solution: Computational prediction of properties in assembling and assembled systems.
Bottleneck/Challenge: Isolation of materials from complex cellular systems.
Potential Solution: Cell-free systems that enable unique protein tagging and purification.
Engineering Biology (2019) Breakthrough Capabilities
Included in the roadmap are select breakthrough capabilities from our 2019 roadmap, Engineering Biology (below in green; milestones at 2021, 2024, 2029, and 2039). While these breakthrough capabilities were written in the context of advancing the field of engineering biology, the EBRC Materials Roadmapping Working Group leading this roadmapping project felt that the technical achievements elaborated in these breakthrough capabilities and their milestones directly contribute to achieving advancements in materials from engineering biology. This content has been incorporated as reference and, when pertinent, will be provided with context for its inclusion in this roadmap.
PCR, reverse transcription, cellular replication, and transcription of fully unnatural nucleotide-containing genes of up to 400 base pairs.
At this length, unnatural aptamer and aptazyme polymers could be regularly evolved and engineered.
Identification of “missing” functionality or functionalities in A-T-G-C base pairs.
Bottleneck/Challenge: Previous work in this field has focused on achieving unnatural base pair incorporation rather than on the incorporation of “useful” bases with specialized chemical functionalities in mind (e.g., metal chelators, novel functional groups etc.).
Potential Solution: Potentially useful chemical functionalities should be enumerated.
Improved in vitro manipulation of unnatural nucleic acids.
Bottleneck/Challenge: Evolution of unnatural aptamers, allosteric regulators and aptazymes requires reverse transcription in order to complete cycles of synthesis and selection.
Potential Solution: Evolve/engineer reverse transcriptases that can incorporate the array of unnatural nucleotides and be able to be easily adjusted to incorporate additional chemistries as they are developed.
Expansion of unnatural nucleotide toolkit.
Bottleneck/Challenge: At present, transcription and translation of DNA containing unnatural base pairs has been achieved only in the context of a single specialized unnatural base pair and only in Escherichia coli.
Bottleneck/Challenge: Success in this endeavor required extensive optimization.
Potential Solution: Begin to explore alternative (previously explored) unnatural base pairs in the context of the optimized conditions, especially ones that do not perturb the double helical structure of DNA and can be incorporated in any sequence context.
Biosynthesis of unnatural nucleotides.
Bottleneck/Challenge: Current unnatural base pairs must be chemically synthesized, which could limit the ability to use them in large scale applications.
Potential Solution: Engineered biosynthetic pathways capable of generating non-natural bases in vivo.
Bottleneck/Challenge: Considerations must be made regarding the risks and impacts of release of non-natural nucleic acids into nature.
Organisms capable of full replication, maintenance, and transcription of a plasmid or artificial chromosome made up entirely of unnatural bases.
Bottleneck/Challenge: Transcripts should confer useful function for the organism and also be made entirely of unnatural bases.
Expanded genetic code systems for translation of >100-amino acid proteins containing fully-unnatural amino acids, and proteins with at least four, distinct unnatural amino acid building blocks.
Implementing unnatural amino acids for materials synthesis will require advancements in translation system engineering, to enable a broader production range from ribosomes. Furthermore, biosynthesis of sequence-defined synthetic biopolymers in which new chemistries (synthetic amino acids or synthetic monomers made-up of non-natural backbones) can be encoded in a template-directed manner, is likely to require the engineering or repurposing of organisms with multiple open coding channels (recoded genomes).
Create proteins that are capable of gaining new, therapeutically-useful activities through unnatural amino acids.
Bottleneck/Challenge: Efficiency and scale of protein expression with expanded genetic code systems.
Potential Solution: Specialized strains, specialized tRNA/aminoacyl-tRNA synthetase systems, specialized ribosomes, and genomically-recoded organisms.
Efficient biosynthesis of proteins containing three or more distinct unnatural amino acid building blocks.
Bottleneck/Challenge: Not enough free codons to hijack for unnatural amino acid building blocks.
Potential Solution: Genomically-recoded strains that free up redundant codons, orthogonal ribosomes that can utilize special tRNAs for special messages, or orthogonal organellar genetic codes.
Bottleneck/Challenge: Not enough mutually orthogonal aminoacyl-tRNA synthetase (aaRS)/tRNA pairs for unnatural building block incorporation.
Potential Solution: Design or scalable evolution of new mutually orthogonal sets of aaRS/tRNA pairs for genomically-recoded organisms or for orthogonal ribosomes; hijacking of organellar genetic codes and associated aaRS/tRNA pairs.
Biosynthesis of unnatural amino acids.
Bottleneck/Challenge: Current unnatural amino acids are chemically synthesized in most cases, which could limit the ability to use them in large scale applications.
Potential Solution: Engineered biosynthetic pathways capable of generating unnatural amino acids in vivo.
Templated biosynthesis and evolution of new polymers with large user-selected sets of unnatural building blocks in vivo.
Bottleneck/Challenge: Ribosome, elongation factor, and aaRS/tRNA engineering for new building blocks and polymer linkage chemistries beyond peptide bonds.
Ability to rationally engineer sensor suites, genetic circuits, metabolic pathways, signaling cascades, and cell differentiation pathways.
The engineering of circuits and pathways will be necessary for the synthesis of material components from engineered biology. The engineering of sensor suites and signalling cascades will also be important for the dynamic behaviors of materials, particularly living materials and composite materials that incorporate cells. More on engineering dynamic activities of materials can be found in Properties & Performance.
Reliable engineering of genetic circuits with more than ten regulators for sophisticated computations.
Bottleneck/Challenge: The availability of orthogonal, programmable, non-repetitive regulators with desired gene regulatory effects.
Potential Solution: Toolboxes of highly non-repetitive, regulated promoters, including large sets of mutually orthogonal promoter/regulator pairs.
Potential Solution: Toolboxes of highly non-repetitive transcription factors or CRISPR genetic parts, including mutually orthogonal transcription factors and CRISPR systems.
Potential Solution: Toolboxes of highly non-repetitive RNA regulators of gene expression that can control transcription, translation and mRNA degradation.
Potential Solution: Toolboxes of highly non-repetitive and mutually orthogonal recombinases.
Potential Solution: Predictive models at nucleotide to systems resolution, capable of predicting how combinations of genetic parts lead to desired computations (e.g., analog, digital, signal processing, pattern recognition).
Reliable engineering of novel, many-enzyme pathways utilizing combinations of bioprospected enzymes with well-characterized kinetics.
Bottleneck/Challenge: Predicting enzymatic reactions and kinetics from amino acid/nucleic acid sequence remains difficult. Many protein or RNA-based enzymes are promiscuous with varying substrate selectivities.
Potential Solution: The kinetics and substrate selectivities of large families of bio-prospected enzymes could be characterized to identify rules for quantifying enzyme-substrate promiscuity.
Potential Solution: Enzyme databases could be greatly expanded to incorporate predicted enzyme-substrate promiscuities, enabling the design of more novel and exotic multi-enzyme pathways.
Five-time improvement and expansion of inducers/promoters for model organisms that respond to environmental inputs and any intracellular metabolite.
Bottleneck/Challenge: We are still relatively limited to chemical inducers of expression.
Potential Solution: Catalog known signals that microbes respond to including potential genetic parts necessary.
Potential Solution: Expand technology to include other non-conventional inputs (e.g., light, electricity); two-year goals would include focus on improving known inputs (e.g., optogenetic tools) and conducting initial tests with new technologies.
Potential Solution: Develop RNA and protein biosensors that respond to a variety of non-natural chemical inputs.
Utilize machine-learning approaches to use the vast amount of uncurated literature results within pathway design.
Bottleneck/Challenge: The performance of pathway design software critically dependent on the quality and amount of enzymatic and biochemical data available; although curated databases such as MetaCyc or BRENDA exist for enzymes involved in metabolism, a large amount of previous experimental results are “buried” in the literature, including records of enzyme specificity and other characterizations, and therefore impossible to utilize for pathway design.
Potential Solution: Develop natural language processing (NLP) and machine learning approaches to extract and characterize the relevant information from the literature of the last four decades.
Creation of optogenetic tools for in vivo RNA post-transcriptional control to allow for easy control of any gene expression process through mRNA.
For reference, please see: You, M., & Jaffrey, S. R. (2015). Designing optogenetically controlled RNA for regulating biological systems.
Annals of the New York Academy of Sciences, 1352, 13–19.
View publication.
Bottleneck/Challenge: There are currently no optogenetic RNA aptamers that can be used in vivo.
Potential Solution: The development of light activated chromophore ligands that are compatible with cellular biochemistry and can be easily synthesized by the cell.
Potential Solution: The design or evolution of aptamers that bind to only one conformation of the chromophore, thus giving the basis for optogenetic switching of RNA structure.
Potential Solution: Enzymatic processes to covalently link RNA sequences to chromophores in a selective manner so that optogenetic chromophore transitions can be cycled as is the case with optogenetic protein mechanisms.
Potential Solution: Take advantage of emerging optogenetic variants of CRISPR/Cas systems, now ported (via Cas13-like effectors) to bind to RNA with light-controlled affinity.
Bottleneck/Challenge: There are currently no optogenetic RNA regulatory mechanisms that can be used in vivo.
Potential Solution: Couple the optogenetic aptamer system to RNA regulatory mechanisms that can control a range of gene expression processes.
Reliable expression of redesigned synthases to produce secondary metabolites, including polyketides and non-ribosomal peptides.
Bottleneck/Challenge: Many important natural products are synthesized by modular multi-enzyme synthases but these complexes are difficult to express, particularly so in engineered forms where specific modules are recombined/designed in order to alter productive specificity.
Potential Solution: Structural analysis, driven by advances in cryo-EM, to inform structural modeling of hierarchical assembly rules.
Potential Solution: High-throughput assembly/assay of natural polyketide synthases (PKSs) and non-ribosomal peptides (NRPs) to build functional PKSs/NRPSs that produce unnatural molecules and incorporate successes/failures into model.
Computational design of protein-ligand and RNA-ligand interfaces suitable for engineering protein-based or RNA-based sensors.
Bottleneck/Challenge: Potential energy models for nucleic acid interactions require improvements in charge screening and ionic effects.
Bottleneck/Challenge: The number of RNA-ligand interactions that have been characterized at atomic resolution is sparse, prohibiting informatics-based approaches to design RNA-ligand interactions.
Potential Solution: Develop and apply high throughput methods to characterize RNA-ligand interactions.
Bottleneck/Challenge: Many candidate RNA/protein sequences must be experimentally characterized to find one that binds well to a targeted ligand.
Potential Solution: Maximally informative measurements of designed proteins and RNAs could be utilized to further improve potential energy functions and model predictions.
Simultaneous, tunable, timed expression of many transcription factors controlling mammalian cell state.
Bottleneck/Challenge: Insufficient mapping between transcription factor expression levels, regulated promoter activities, epigenetic modifications, and overall regulation of protein and metabolite levels.
Bottleneck/Challenge: Insufficient temporal expression control over multi-protein genetic systems in eukaryotic systems, particularly mammalian systems.
Potential Solution: Development of improved genetic circuits controlling temporal expression of desired proteins in mammalian cells, stimulated by small molecule, cell contact, or cell cycle conditions.
Potential Solution: Systematic characterization of human, genome-wide transcription factor binding sites, affinities, and gene regulatory effects.
Ability to build and control small molecule biosynthesis inside cells by design or through evolution.
In the context of synthesizing materials from engineering biology, design and evolution of cells to build and control the synthesis of monomers, in addition to small molecules, will be important.
Identify model organisms for performing specific types of chemistries or organisms that have native precursor biosynthesis pathways for specific classes of molecules.
Bottleneck/Challenge: Academics and companies have constructed heterologous hosts for a limited number of chemical classes, but there are many other chemical classes that need hosts with precursor pathways constructed; additionally, some of these precursor production hosts may not be ideal for particular applications or environments.
Potential Solution: Use bioinformatics and screening to identify organisms that might be particularly useful for producing a chemical under a particular environmental condition or that already has precursor pathways for a particular class of chemicals; build a database of those organisms.
Potential Solution: Development of robust plant hosts in order to bridge the gap before trying to engineer plant metabolic pathways into microbes.
Precise temporal control of gene expression for well-studied systems.
Bottleneck/Challenge: Many desired chemicals are toxic to growth of the producing host. Avoidance of this results in separation of growth and production phases which has been achieved by changing the media conditions; however, such approaches are costly, may require the introduction of extra chemicals that are difficult to remove from the desired products, and cannot easily accommodate cellular heterogeneity, or be used to fine tune the shift from growth to production phases.
Potential Solution: Develop several types of expression systems for controlling the timing of gene expression and thus the timing of chemical production; ultimately, decouple growth and reproduction from energy and carbon metabolism and product generation.
Bottleneck/Challenge: The necessary genetic regulatory elements should be sufficiently orthogonal to the host to permit tuning, but nonetheless responsive to changes in cell state.
Potential Solution: Identifying, and then re-engineering, diverse sets of protein and RNA regulators capable of activating or repressing gene expression in response to targeted changes in growth and nutrient state would expand these capabilities; integrating the responses of these regulatory elements, perhaps through the use of engineered information processing networks, would permit the construction of circuitry for fine-tuning the timing of gene expression in microbial hosts.
Construct a limited number of model host organisms for synthesizing all-natural products.
Bottleneck/Challenge: There is a need for hosts capable of producing all natural products found in nature; there is no need for a single host to produce all of the natural products, but rather at least one host for every class of natural products.
Potential Solution: Use the database of possible hosts, genetic control systems and modular pathways to develop a range of hosts for each class of natural products; these hosts should produce high levels of the precursors to natural products.
Construction of single-cell organisms for production of unnatural derivatives of natural products.
Bottleneck/Challenge: There are few engineered cells that produce unnatural derivatives of natural products and the methods for engineering organisms to produce unnatural natural products are nascent.
Potential Solution: Construct hosts with the modular natural precursor pathways and engineered or evolved enzymes to create hosts that can produce unnatural natural products.
Potential Solution: Construct hosts with promiscuous enzymes (either natural, engineered, or evolved enzymers) and feed unnatural precursors that are incorporated into the final product.
Temporal control over multiplexed regulation of many genes in parallel.
Bottleneck/Challenge: It remains difficult to target gene expression to a particular growth phase or culture time in order to maximize production of a desired natural or unnatural product.
Potential Solution: Develop a catalog of promoters (including timing and gene expression strength) that are activated in various phases of cultivation.
Potential Solution: Identify promoters that are activated at the same time or with the same environmental queue and that can be used with multiple genes (i.e., multi-step biosynthetic pathway).
Software and hardware for optimizing titer, rate, and yield of any product produced by any host.
Bottleneck/Challenge: It is challenging and time-consuming to optimize the titer, rate and yield of a desired product in any host.
Potential Solution: Develop a software suite that considers the metabolic pathway(s) and host and incorporate a wide variety of measurements and can predict that changes that need to be made to the metabolic pathway add the host to optimize titer, rate, and yield.
On-demand construction of single cell organisms for production of nearly any molecule of interest, including organic chemicals and polymers.
Bottleneck/Challenge: Reliance on nature’s single-cell organisms that are not ideally suited for producing a particular chemical.
Potential Solution: Use retrobiosynthesis software as well as genetic control system design software to de novo design microbial cells that can produce nearly any organic chemical; the cells would be designed for the processing environment to enable inexpensive purification of the final product.
Ability to manufacture any targeted glycosylated protein or metabolite using cell-free biosynthesis.
Contributors to this roadmap have indicated that the 2024 milestone “Production of bacterial glycoconjugate vaccines in cell-free systems” would gain significant relevance and value by including the production of bacterial glycoconjugate therapeutic antibodies in cell-free systems, in addition to vaccines.
Ability to build modular, versatile cell-free platforms for glycosylation pathway assembly.
Bottleneck/Challenge: Many of the most important components of glycosylation pathways are associated with cellular membranes and cannot be recapitulated easily in cell-free systems.
Potential Solution: Develop and optimize efficient strategies use of oligosaccharyltransferases (including eukaryotic versions) to transfer pre-built sugars from lipid-linked oligosaccharides in vitro.
Potential Solution: Develop methods to select and assemble a set of soluble enzymatic tools capable of producing therapeutically-relevant glycoproteins with desired properties in vitro from simple, commercially available activated-sugar building blocks.
Expanded set of glycosylation enzyme-variants that efficiently install eukaryotic glycans
Bottleneck/Challenge: Synthesis of complex human glycans in cell-free systems is constrained by the available set of well characterized enzymes; existing characterizations do not provide sufficient tools to go from a set of sugar monomers and a design to a glycoprotein of interest.
Potential Solution: Expand the glycoengineering toolkit by characterizing glycoslytransferases to assemble sets of well-characterized enzymes that can reliably produce desired glycoproteins.
Potential Solution: Engineer existing glycosylation enzymes for desired activities when naturally occurring enzymes with desired functionalities are not available (e.g., engineering of the bacterial oligosaccharyltransferase to accept the eukaryotic core glycan would enable the efficient production of the eukaryotic core glycan in bacterial systems).
Production of bacterial glycoconjugate vaccines in cell-free systems.
Bottleneck/Challenge: The production of glycoconjugate vaccines against bacteria require the culturing of pathogenic strains recalcitrant to bioengineering and the use of non-specific conjugation chemistry.
Potential Solution: Express diverse bacterial O-antigen pathways for characterization and production of lipid-linked oligosaccharides required for vaccines and use bacterial oligosaccharyltransferases to site-specifically attach these oligosaccharides in vitro.
Potential Solution: Use the greater control afforded by cell-free protein synthesis systems to produce FDA-approved vaccine carrier and antigen proteins, many of which cannot be produced outside of pathogenic organisms.
Expanded set of enzymes capable of glycosylating metabolites in vitro.
Bottleneck/Challenge: Glycosylation of therapeutically-relevant metabolites (including many antibiotics) is often required for desirable pharmacokinetic/dynamic properties, but many of the enzymes which perform these glycosylation activities are unknown or can only be expressed in their native host strain.
Potential Solution: Use cell-free protein synthesis to screen the activity of metabolite-targeting glycosyltransferases on existing bioactive compound libraries to understand their specificities and improve the therapeutic utility of small molecule products.
Bottleneck/Challenge: Substrate limitations lead to inherent inefficiencies.
Potential Solution: Develop metabolic models of lysate hosts to identify genetic knockouts that enhance glycan production and carry out such modifications.
Cell-free pipelines to produce and assess the functionality of diverse, human glycosylated protein therapeutics.
Bottleneck/Challenge: Development timelines of glycoprotein therapeutics are slowed by the need to produce these products in mammalian cell lines.
Potential Solution: Develop, optimize, and implement strategies to create more than ten unique and homogeneous glycan structures on proteins by cell-free methods (e.g., trimannose core eukaryotic glycan, afucosylated galactose-terminated core glycan, fucosylated sialic-acid terminated core glycan, biantennary glycan, etc.); once synthesized, these products can studied and optimized using therapeutic functionality assays.
Ability to produce any glycosylated protein therapeutics and vaccines at the point-of-care in less than one week.
Bottleneck/Challenge: Intentional engineering of glycan structures and the synthesis of novel structures is constrained by identifying sets of enzymes for the manufacture of the sugar structures.
Potential Solution: Develop cell-free systems capable of rapidly and robustly producing any defined glycoprotein on-demand.
Production and secretion of any protein with the desired glycosylation or other post-translational modifications.
In the context of synthesizing materials from engineering biology, this breakthrough capability can include the production and secretion of any small polypeptide, in addition to protein, with the desired modifications, and non-ribosomal protein production. Post-translational modifications that are likely to greatly contribute to protein and polypeptide synthesis for materials include phosphorylation, acetylation, and methylation, among other less-common modifications.
One or more microbial hosts capable of producing laboratory-scale quantities of a single glycoform of a desired protein.
Bottleneck/Challenge: Compared to protein synthesis, post-translational modification and machinery is not well understood and any given protein may exist in the cell in multiple glycoforms, making advanced study challenging; this problem is particularly prevalent in the production of biologic pharmaceuticals (e.g., biosimilars) where the complex array of glycosylation patterns can cause differences in efficacy and safety.
Potential Solution: Understand the pathways involved in glycosylation of the protein of interest and develop these pathways to enable tightly controlled synthesis of a single glycoform.
A few microbial hosts capable of secreting functional versions of proteins with no post-translational modifications.
Bottleneck/Challenge: There are relatively few bacterial hosts that have well developed protein secretion systems. Those that exist are primarily in Gram positive hosts, and are not necessarily compatible with all desired protein products; gram negative hosts, on the other hand, have an outer membrane that must also be crossed before exiting the cell, and secreting proteins across both the inner and outer membranes remains a challenge at the efficiencies required for commercial application.
Potential Solution: Develop new bacterial hosts with machinery specifically designed for protein secretion.
Bottleneck/Challenge: The systems that do exist are either required for cellular function and thus can only be repurposed to a limited extent, or are highly regulated and difficult to activate or keep activated.
Potential Solution: Utilize systems biology to understand and remove the regulatory controls on secretion systems.
Ubiquitous control of post-translational modification (including glycosylation of multiple sites with multiple sugars) in a diverse array of hosts.
Bottleneck/Challenge: Post-translational modifications are often considered the “analog” control in the cell; as such, they can carefully tune how an individual protein interacts with its environment. The lack of understanding of this control makes affecting post-translational modification for desired use particularly challenging. The ability to tune the analog portion of the cell, however, can allow for more efficient systems, better synthetic yields, and a broader set of uses than would otherwise be possible.
Long-lasting, robust, and low-cost cell-free system for protein synthesis and biomanufacturing.
Identify reagent instabilities in cell-free systems across multiple organisms and all biological kingdoms.
Bottleneck/Challenge: Stability of reagents is not well understood in non-E. coli-based cell-free systems.
Potential Solution: The full spectrum of metabolic activity during cell-free reactions over time can be characterized in multiple systems, including those emerging cell-free systems, to identify reagent instabilities.
Alleviate reagent instabilities and prolong the half-life of cell-free reagents from a few hours to several days using inexpensive substrates.
Bottleneck/Challenge: High-energy phosphate compounds are still commonly used to fuel cell-free systems (e.g., PEP, 3PGA); these compounds are expensive, lead to inhibitory phosphate concentrations, and only enable bursts of ATP regeneration instead of long-lived energy regeneration.
Potential Solution: Central metabolism, non-phosphorylated energy substrates, or light driven approaches (e.g., from bacteriorhodopsin), among others, could be used to enable long-lived energy regeneration without inhibitory byproducts; should be tested in extracts made from organism across all kingdoms and identify key factors (such as, concentrations, additional reagents, etc.) for robust alternative energy activation.
Bottleneck/Challenge: Cofactor regeneration and balancing strategies preclude long-term activation of energy metabolism to fuel metabolic activity.
Potential Solution: Molecular purge valves implemented in crude extracts could maintain redox balance.
Potential Solution: Develop schemes for balancing ATP, and derivatives including NAD(P) and FAD.
Potential Solution: Create non-natural cofactors, and requisite engineered enzymes, that have better stability properties.
Bottleneck/Challenge: Reagent instabilities limit reaction time.
Potential Solution: Genomic modifications to extract source strains can be carried out to stabilize substrates.
Avoid inhibition (poisoning) of cell-free reactions by byproducts or the desired products.
Bottleneck/Challenge: Small molecule byproducts of metabolism, such as phosphate, inhibit cell-free reactions.
Potential Solution: Engineer systems to avoid the accumulation of chemical inhibitors.
Potential Solution: Develop product siphoning strategies and dialysis-like strategies to remove inhibitors from the system.
Potential Solution: Engineer molecular complexes that are resistant to byproducts and chemical products.
Stabilize catalysts to facilitate cell-free reactions on the order of weeks.
Bottleneck/Challenge: Cell-free reactions terminate because substrates and cofactors are depleted, byproducts accumulate to inhibit the reaction, or the catalysts become inactivated; of these, enzyme stability represents a significant technical and economic hurdle to technology development in this space.
Potential Solution: Characterize catalyst instabilities such as, tolerance to byproducts and inhibitors.
Potential Solution: Increase stability of enzymes involved in cell-free systems.
Potential Solution: Generate design criteria for choosing catalysts (i.e., identifying select organisms from which to derive lysates and enzymes) for desired applications.
Potential Solution: Develop reactor designs and bioprocesses that continuously replenish the source of catalysts.
Robust and scalable production of cell-free systems that last for weeks.
Bottleneck/Challenge: Without enzymes or lysates that are stable on the order of weeks, significant fractions of carbon will otherwise be used in generating the biocatalysts required of these systems.
Potential Solution: Integrate knowledge of the system and innovations to facilitate long-lived reactions.
Footnotes
- Arranz-Gibert, P., Vanderschuren, K., & Isaacs, F.J. (2018). Next-generation genetic code expansion. Current Opinion in Chemical Biology, 46, 203-211. https://doi.org/10.1016/j.cbpa.2018.07.020
- Lutz, J-F., Ouchi, M., Liu, D.R., & Sawamoto, M. (2013). Sequence-controlled polymers. Science, 341(6146), e1238149. https://doi.org/10.1126/science.1238149
- Thuronyi, B.W., Privalsky T.M., & Chang, M.C.Y. (2017). Engineered fluorine metabolism and fluoropolymer production in living cells. Angewandte Chemie International Edition, 56(44), 13637-13640. https://doi.org/10.1002/anie.201706696
- Panganiban, B., Qiao, B., Jiang, T., DelRe, C., Obadia, M.M., Nguyen, T.D., Smith, A.A.A., Hall, A., Sit, I., Crosby, M.G., Dennis, P.B., Drockenmuller, E., Olvera de la Cruz, M., & Xu, Ting. (2018). Random heteropolymers preserve protein function in foreign environments. Science, 359(6381), 1239-1243. https://doi.org/10.1126/science.aao0335
- Ribosome pilot challenge. Eterna. https://eternagame.org/labs/9162726 (Accessed January 6, 2021.)
- Hammerling, M.J., Fritz, B.R., Yoesep, D.J., Kim, D.S., Carlson, E.D., & Jewett, M.C. (2020). In vitro ribosome synthesis and evolution through ribosome display. Nature Communications, 11(1108). https://doi.org/10.1038/s41467-020-14705-2
- Lajoie, M.J., Rovner, A.J., Goodman, D.B., Aerni, H.R., Haimovich, A.D., Kuznetsov, G., Mercer, J.A., Wang, H.H., Carr, P.A., Mosberg, J.A., Rohland, N., Schultz, P.G., Jacobson, J.M., Rinehart, J., Church, G.M., & Isaacs, F.J. (2013). Genomically recoded organisms expand biological functions. Science, 342(6156), 357-360. https://doi.org/10.1126/science.1241459
- Johnston, T.G., Yuan, S., Wagner, J.M., Johnston, T.G., Yuan, S., Wagner, J.M., Yi, X., Saha, A., Smith, P., Nelson, A., & Alper, H.S. (2020). Compartmentalized microbes and co-cultures in hydrogels for on-demand bioproduction and preservation. Nature Communications, 11, 563. https://doi.org/10.1038/s41467-020-14371-4
- Amiram, M., Haimovich, A.D., Fan, C., Wang, Y., Aerni, H., Ntai, I., Moonan, D.W., Ma, N.J., Rovner, A.J., Hong, S.H., Kelleher, N.L., Goodman, A.L., Jewett, M.C., Söll, D., Rinehart, J., & Isaacs, F.J. (2015). Evolution of translation machinery in recoded bacteria enables multi-site incorporation of nonstandard amino acids. Nature Biotechnology, 33, 1272–1279. https://doi.org/10.1038/nbt.3372
- Vermaas, J.V., Dellon, L.D., Broadbelt, L.J., Beckham, G.T., & Crowley, M.F. (2019). Automated transformation of lignin topologies into atomic structures with LigninBuilder. ACS Sustainable Chemistry & Engineering, 7(3), 3443–3453. https://doi.org/10.1021/acssuschemeng.8b05665
- Hadadi, N. & Hatzimanikatis, V. (2015). Design of computational retrobiosynthesis tools for the design of de novo synthetic pathways. Current Opinion in Chemical Biology, 28, 99-104. https://doi.org/10.1016/j.cbpa.2015.06.025
- Caspi, R., Billington, R., Fulcher, C. A., Keseler, I. M., Kothari, A., Krummenacker, M., … Karp, P. D. (2018). The MetaCyc database of metabolic pathways and enzymes. Nucleic Acids Research, 46(D1), D633–D639. https://doi.org/10.1093/nar/gkx935 MetaCyc is available at https://metacyc.org.
- Jeske, L., Placzek, S., Schomburg, I., Chang, A., & Schomburg, D. (2019). BRENDA in 2019: a European ELIXIR core data resource. Nucleic Acids Research, 47(D1), D542–D549. https://doi.org/10.1093/nar/gky1048 BRENDA is available at https://www.brenda-enzymes.org/.
- Clark, D. S., & Blanch, H. W. (1997). Biochemical Engineering (Chemical Industries) (2nd ed., p. 716). New York, New York: Crc Press.
- Venayak, N., von Kamp, A., Klamt, S., & Mahadevan, R. (2018). MoVE identifies metabolic valves to switch between phenotypic states. Nature Communications, 9(1), 5332. https://doi.org/10.1038/s41467-018-07719-4
- Weinhandl, K., Winkler, M., Glieder, A., & Camattari, A. (2014). Carbon source dependent promoters in yeasts. Microbial Cell Factories, 13, 5. https://doi.org/10.1186/1475-2859-13-5; Hsiao, V., Cheng, A., & Murray, R. M. (2016). Design and application of stationary phase combinatorial promoters (SEED 2016 Technical Report). Retrieved from http://www.cds.caltech.edu/~murray/preprints/hcm16-seed_s.pdf
- National Research Council (US) Committee on Assessing the Importance and Impact of Glycomics and Glycosciences. (2012). Transforming glycoscience: A roadmap for the future. Washington (DC): National Academies Press (US). https://doi.org/10.17226/13446
- Sethuraman, N., & Stadheim, T. A. (2006). Challenges in therapeutic glycoprotein production. Current Opinion in Biotechnology, 17(4), 341–346. https://doi.org/10.1016/j.copbio.2006.06.010
Last updated: January 19, 2021
