Engineering Biology & Materials Science
Processing
Processing considers the engineering of biology to conduct “unit operations” to build or destroy materials through polymerization and degradation, templating, patterning, and printing.
Introduction
The Processing technical theme considers advancements in engineering of biology to conduct “unit operations” to build or destroy materials through polymerization and degradation, templating, patterning, and printing. This includes engineering the biological extrusion or secretion of materials, material deposition, and self-assembly and -disassembly. Processing also includes engineering biology-based technologies, tools and systems (e.g., cell-free systems) to manufacture, recover, and purify materials. Includes engineering biological materials to function in non-natural environments and extreme conditions.
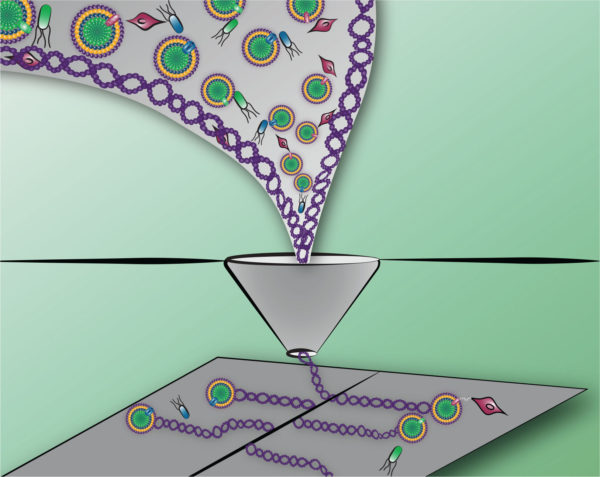
Advancements in processing of materials with, or directed by, biocomponents can enable new characteristics and functionalities. Depicted here, synthetic polymers with differing dimensionality are sorted and printed in a manner that localizes cells on a surface. The complex patterning of bioactive material is enabled by polymers that provide protection through a printing process. Processing of materials systems (e.g., extracellular matrix mimetics) in a manner that can distinguish and sort various components of the system for precise deposition into patterns provides a degree of control and reproducibility not available in natural systems.
Processing Breakthrough Capabilities
Enable secretion of monomers or polymers without destruction of cells.
Enable efficient protein/polypeptide secretion, including proteins containing non-standard amino acids.
Bottleneck/Challenge: Secretion systems in model organisms are limited in yield and sequence requirements.
Potential Solution: Develop better tools for protein secretion and for polymer formation in non-model organisms (e.g., Bacillus subtilis).
Bottleneck/Challenge: Limited number of chassis used for non-standard amino acid incorporation.
Potential Solution: Expand orthogonal translational systems to other organisms such as Bacillus subtilis, an efficient protein secretor.
Advance bioprocess techniques used in fermentation to isolate secreted hydrophobic materials.
Bottleneck/Challenge: Limited secretion products have resulted in under-exploration of in situ separation methods to improve production.
Potential Solution: Test techniques such as organic overlay, used for in situ small molecule extraction.
One-pot fermentation and conjugation or macromolecular assembly for streamlined separation.
Bottleneck/Challenge: Few conjugation, functionalization, and/or assembly strategies well-suited to occur in or around cells.
Potential Solution: Identify or design new coupling chemistries that function in fermentative environments.
Design and engineer eukaryotic chassis and secretion systems that allow compartmentalization and timing of synthetic pathways, taking inspiration from developmental biology.
Bottleneck/Challenge: Incomplete understanding of, and design tools, for complex regulation that controls spatiotemporal patterning in biological systems.
Potential Solution: Identify a series of model materials with design features that require spatiotemporal patterning and compartmentalization, to quantify progress.
Incorporate coordinated consortia of production chassis that can build (in situ) complex patterned materials from diverse unit-level materials (i.e., diatom deposition of silica combined with bacterial protein functionalization).
Bottleneck/Challenge: Lack of cooperativity in growth and material production and patterning in organisms that do not come by it naturally; inconsistent growth medium requirements and replication rates that confound such coordinated consortia.
Potential Solution: Engineer mutualistic dependencies between disparate organisms that link growth and development, providing the framework on which to build spatiotemporal coordination biomaterial synthesis and deposition.
Potential Solution: Template inorganic structures with unique stoichiometry and potential to access novel ceramics or catalysts.
Ability to control self-assembly and disassembly of biomolecule-based or -embedded materials.
Development of materials design approach that allows for programmatic control of self-assembly and disassembly.
Bottleneck/Challenge: Design rules of self-assembly are not fully elucidated.
Potential Solution: Development of experimental and computational frameworks to support materials discovery for biomolecule-based and/or embedded materials (e.g., machine learning, computational biology, and protein-folding algorithms).
Understand the mechanisms of cellular remodeling of the extracellular matrix (ECM) and dynamic interactions between cells and templating materials.
Bottleneck/Challenge: Fundamental gaps in knowledge about how specific protein sequences and tissue inhibitors affect cellular properties and cell-matrix interactions.
Potential Solution: Controlled engineering of three-dimensional networks that resemble the ECM.
Potential Solution: Standardized characterization of cellular properties in natural and engineered networks with site-specific cellular/protein interactions.
Bottleneck/Challenge: Gaps in knowledge remain about how specific physical properties of ECM (stiffness, porosity, viscoelasticity) affect cell functions from different cell types.
Potential Solution: Enable more thorough characterization of ECM physical properties.
Ability to control and direct hierarchical self-assembly to achieve desired material architecture, properties, or functions.
Bottleneck/Challenge: Limited understanding of the complex multiscale interactions that drive self-assembly.
Potential Solution: Molecular-level investigations of the self-assembly of stimuli-responsive biopolymers or amphiphiles with conformational adaptability.
Bottleneck/Challenge: Understanding of the interactions of biocomponents with the substrate, including covalent, ionic, and van der Waals interactions.
Potential Solution: Characterize the bonding in categories of the type of interaction, with the aim that similar biocomponents will bond similarly to the same substrate.
Bottleneck/Challenge: Tunable protein-based stimulus-responsive nanostructures.
Potential Solution: Characterize and engineer existing proteins that are responsive to light, pH, mechanical stimuli, etc.
Ability to control molecular and macromolecular deposition, patterning, and remodelling on biotic and abiotic surfaces.
Increase capability to deposit or pattern biomolecules on various substrates.
Bottleneck/Challenge: Lack adhesive biological and/or synthetic moieties that allow for specific interactions.
Potential Solution: Engineer a series of orthogonal membrane proteins on living cells to create autonomous patterns on surfaces.
Bottleneck/Challenge: Limited characterization methods for patterned/deposited surfaces.
Potential Solution: Establish and expand dimension, thickness, periodicity, and spectroscopic methods.
Control proteins on the outside of an organism for selective reactivity with material precursors for the creation of size and shape controlled materials.
Bottleneck/Challenge: Understanding of amino acid and traditional chemical reactivity to build controlled structures.
Potential Solution: Full characterization of the suites of materials produced to date and analyze with artificial intelligence and machine learning techniques.
Ability to build robust interfaces between living systems and semiconducting/electronic surfaces to sense and manipulate biological processes.
Bottleneck/Challenge: Biological materials do not withstand the processing conditions that semiconductors do.
Potential Solution: Stepwise generation of hybrid systems to account for the fragility of biocomponents.
Potential Solution: Depending on application/desired outcome, inorganic semiconductor may not be needed, and a more biocompatible one, such as an organic semiconductor, may be used with processing conditions closer to that of biomaterials.
Bottleneck/Challenge: Molecular platforms to transmit electrical signals between cells and electronic materials are not universal and maybe difficult to integrate with signal transduction pathways.
Potential Solution: Develop interchangeable tools to allow for integration of common electrical signals (e.g., redox, ion conduction).
Produce templated materials of atypical biological shapes or size scales.
Bottleneck/Challenge: The utilization of what we already know in templating materials on biological systems to produce the material size and shapes desired.
Potential Solution: Utilize techniques such as DNA origami or very selective chemistry to combine biological materials into non-natural shapes and sizes (e.g., combining linear phage into the vertices of a pyramid).
Potential Solution: Manipulate an organism to autonomously form a desired size or shape.
Potential Solution: Use bilayers, micelles, and vesicles to constrain shape.
Potential Solution: Use a patterned and sacrificial substrate to constrain systems to certain shapes.
Potential Solution: Use asymmetric polymeric materials (e.g., tapered bottlebrushes similar to scaffolding proteins) as molecular units for templating and scaffolding.
Engineer living organisms that change their surface properties upon response to an external stimuli, in order to template desired materials on demand.
Bottleneck/Challenge: Multiple biological processes need to work in tandem, molecular precursors need to be present and accessible, and the kinetics of many processes need to be improved.
Potential Solution: Improve engineered communication between singular intracellular functions.
Potential Solution: Perform processes on an intelligent substrate, such as one that can monitor activity of components or precursors and update components’ information to each other.
Controllably form on-demand macroscale materials templated on biological organisms with desired properties.
Bottleneck/Challenge: Controlled interactions between individual materials would be required in order to stack materials in a desired array without outside stimuli.
Potential Solution: Innovation in additive manufacturing (e.g., in situ printing of a defined layer of materials directly onto a living organism).
Enable biomolecule and cellular patterning and printing under diverse conditions.
Develop technology to spatially place single cells at defined locations with high spatial resolution in two dimensions, as the biological pattern for the material.
Bottleneck/Challenge: Maintaining the structure and function of cells in the template.
Potential Solution: ‘Ink-jet printing’ of bacterial inoculum using acoustic liquid-handling.
Potential Solution: ‘Photo-masking’ enabled selection of spatially-resolved surviving organisms from a mixed population using light.
Potential Solution: Imprint lithography to generate a scaffold with spatially differentiated layers or regions through the step of coating these regions with an appropriate chemical.
Establish 3D-printing methods for producing biological (living) composite materials.
Bottleneck/Challenge: Cellular behavior in printing conditions (in shear field, solidification, or polymerization) remains to be explored.
Potential Solution: Develop an analysis flow for systematic studies on the relevant variables (e.g., printing condition and viability, scaffold molecular composition and viability, viability and motility relationships with metabolic activity).
Potential Solution: Develop methodology to assess cellular viability during or shortly following printing.
Bottleneck/Challenge: Engineering microstructures within the printed biological composite materials.
Potential Solution: Enable block-copolymer, shear-induced pre-alignment.
Potential Solution: Introduce acoustic, magnetic, or electric fields into synthesis protocols.
Capability to print microbes and/or grow desired three-dimensional communities from two-dimensional patterns under various environmental conditions.
Bottleneck/Challenge: ’Externally’ defined 3D patterns (i.e., those determined by printing) are not currently reinforced by intracellular genetic circuits that preserve patterning in different environments.
Potential Solution: Introduce asymmetric control of cell growth and reproduction, such that one class of microbes constrains physiology of other community members in a consortium.
Maintain desired microbial community structural organization over time.
Bottleneck/Challenge: Engineering persistence of designed and structured 3D cellular communities over time.
Potential Solution: Control individual strains in a mixed-culture pattern with feedback circuits from environmental sensors to preserve deposited patterns in diverse environmental conditions.
Print cellular structures on demand on any surface.
Bottleneck/Challenge: Need to enable a wider variety of cell types that can be printed.
Potential Solution: Further characterization of multi-species and multicellular communities and structures.
Engineer cells to produce materials in environments optimal for the ex vivo material.
Develop additional polymerization strategies (e.g., peroxidase-catalyzed, reversible or irreversible covalent couplings) for abiotic and non-natural monomers that can occur in aqueous or fermentative environments.
Bottleneck/Challenge: Many of these enzymes/couplings function only in conditions incompatible with protein and cell stability.
Potential Solution: Investigate associated enzyme family members under a broader range of conditions.
Engineer minimal cells for production and processing of biological materials (e.g., chromosome-free bacterial cells).
Bottleneck/Challenge: Expanding types and host chassis for creating minimal cell systems is required.
Potential Solution: Explore types and host chassis for creating minimal cell systems and test their resilience, robustness, and performance in/with materials.
Engineer thermophilic microorganisms for producing biomass or biomolecules to produce materials in/at higher temperatures.
Bottleneck/Challenge: Engineering of non canonical organisms for isolation of novel molecules.
Potential Solution: Determine synthesis chassis for extremophile organisms that allows production of unique molecules.
Manage biological complexity and interactions in experimental model systems to determine the ideal host species for a product.
Bottleneck/Challenge: Building of models with experimental data from a wide-range of organisms.
Potential Solution: Import of useful functionalities from less conventional organisms into other chassis.
Production and processing of biological materials in the absence of water.
Bottleneck/Challenge: Biological structure-function-property relationships are driven by solvophobic/hydrogen bonding interactions; very difficult to translate to other solvent systems.
Potential Solution: Explore polar, non-volatile solvent systems for ability to maintain enzymatic function, structure in non-aqueous environments; initial substrates may include dimethyl sulfoxide, ethylene glycol, and low-molecular-weight polyethylene glycols.
Potential Solution: Develop predictive capability for folding correlated to solvent dielectric, protein analogs.
Enable robust processing of materials using cell-free systems.
Expand cell-free systems to different bacterial species and mammalian cell types.
Bottleneck/Challenge: Standardization of cell lysates collection and optimization of reaction conditions.
Potential Solution: Systems biology approaches (e.g., proteomics, metabolomics, transcriptomics) to analyze cell-free expression systems to assess and identify limiting factors.
Use cell-free systems for large scale production of complex biologics including proteins with post-translational modifications.
Bottleneck/Challenge: Large-scale culture may require industry scale equipment not readily available in academic labs.
Potential Solution: Develop continuous bioprocessing strategies for routine, large-scale production of cell-free systems.
Bottleneck/Challenge: Post-translational modifications are difficult to precisely control.
Potential Solution: Engineer systems in which cell-free synthesized proteins go through a series of desired modifications by passing through microfluidic channels with enzymes at specific locations to modify the proteins passing through, akin to a conveyer belt in a manufacturing process.
Potential Solution: Develop batch processes with multiple distinct enzymes together that enables synthesis of homogeneous glycoproteins.
Establish cell-free “distributed processing” for sustainable feedstock utilization.
Bottleneck/Challenge: Compartmentalization is typically lacking in cell-free systems.
Potential Solution: Engineer encapsulation into cell-free systems or minimal cells to permit material separation from enzymes or cofactors for optimized utilization.
Augment materials with freeze-dried cell-free systems for on demand, real-time diagnostics.
Bottleneck/Challenge: Humidity and reaction stability limit use.
Potential Solution: Develop abiotic materials that can maintain biological conditions.
Establish cell-free systems for on-demand and personalized biomanufacturing platforms.
Bottleneck/Challenge: High costs limit adoption.
Potential Solution: Develop strategies based on energy substrates and strain engineering to reduce costs an order of magnitude from present day.
Bottleneck/Challenge: Reaction stability constrains utility in resource limited settings.
Potential Solution: Develop materials that stabilize cell-free systems for extended periods of time while freeze-dried (>1 year).
Develop carbon-optimized cell-free bioconversions for the industrial scale production of materials.
Bottleneck/Challenge: Most bioprocesses still use reduced-carbon compounds like glucose.
Potential Solution: Capture carbon from CO2 fixation pathways.
Potential Solution: Combine electrolyzers with cell-free systems to fuel cost-effective, manufacturing strategies at large-scale.
Bottleneck/Challenge: Co-factor regeneration and stability is a concern.
Potential Solution: Develop new stable cofactors.
Bottleneck/Challenge: Enzyme stability is limiting.
Potential Solution: Develop continuous bioprocesses to replenish catalysts.
Potential Solution: Develop materials to stabilize catalysts for one month continuous operation.
Enable selective component and material degradation through engineering biology.
Identify and engineer enzymes (e.g., esterase) to interact and degrade polymeric substrates with tunable performance.
Bottleneck/Challenge: Studies of enzymatic degradation of polymeric scaffolds exist, yet their performance and substrate scope remains to be improved and engineered for various contexts; current approaches involve trial and error and oftentimes substrates are not selective but instead responsive to a variety of enzymes.
Potential Solution: Develop predictive models that help develop structure-property relationships in enzymatically degradable systems that allow for development of materials with increased selectivity.
Identify enzymes capable of interaction with, and degradation of, both hard and soft segments of polymer systems.
Bottleneck/Challenge: Complete degradation of polymers (e.g., ester and carbonyl bonds) may require multi-enzyme systems or chimeric enzymes.
Potential Solution: Enable engineering of expanded classes of enzymes, including incorporation of non-standard and/or non-natural amino acids.
Design and engineer cellular half-life and patterning for persistence in materials.
Bottleneck/Challenge: Spore-forming microbes and associated tools have been underdeveloped for synthetic biology applications.
Potential Solution: Engineer spore-display and genetic circuit tools for responsive functionalization of spores to abiotic interfaces or within abiotic materials.
Engineer enzymes capable of degrading or reversing click chemistry reactions used to conjugate proteins and polymers, while leaving the polymer and protein intact.
Bottleneck/Challenge: Click chemistries used for bioconjugation through the use of non-standard amino acids are infrequent/nonexistent in nature, enzymes that can specifically degrade/reverse a click reaction have not been identified or developed.
Potential Solution: Scour sequence databases for DNA sequences that could encode enzymes with needed putative activities.
Potential Solution: Pursue a protein engineering effort to generate an enzyme capable of degrading or reversing a click reaction.
Engineer living cells to produce and secrete the target enzymes for modulating polymeric scaffold in situ.
Bottleneck/Challenge: Requires development of feedback loops for new materials.
Potential Solution: Studies to explore dynamic equilibria exhibited by actin filaments, application to motility and force generation; develop principles for governing dynamics, required kinetics, expand from baseline understanding.
Enable force application and specific ECM binding (‘AND gate’) to trigger ECM degradation.
Bottleneck/Challenge: Control of sense-response activity for extracellular matrices.
Potential Solution: Identification of signaling nodes that receive input from force sensing (i.e., via a mechanosensitive channel) and ECM binding that can be coupled to secretion of matrix metalloproteinases.
Enable aqueous biomolecules to interact at high affinity with existing insoluble polymers.
Bottleneck/Challenge: Degradative enzymes are often intolerant of solvents required to solubilize synthetic polymers.
Potential Solution: Design strategies to functionalize enzyme surfaces for increased solvent tolerance (e.g., via non-standard amino acids such as fluoro-phenylalanine, post-translational modifications, or synthetic modification).
Potential Solution: Enable a co-polymer approach to enzyme stabilization.
Engineer polyspecific enzymes that can break specific classes of chemical bonds between heteroatoms (C-O, C-N, etc.), also potentially targeting unsaturation (e.g., metathesis).
Industrial infrastructure and accelerated downstream processing of biocomponent-containing materials.
Enable systems that operate in semi-continuous modes with modular units designed using process-intensification principles to accelerate transport and reaction processes.
Bottleneck/Challenge: Efficient product isolation and purification requires biocatalysts compatible with advanced materials and systems engineering strategies spanning multiple scales.
Potential Solution: Bioreactors with cells suspended in extruded pluronic hydrogels with tunable rates of substrate and product diffusion.
Potential Solution: 3D-printed biocompatible nanostructures encapsulating engineered cell- or cell-free biocatalysts.
Establishment of multiple (5+) pilot scale facilities (300-10,000 L) for the production of biomolecules and biomaterials; such facilities each need to enable fermentation, downstream processing, and feedstock supply.
Bottleneck/Challenge: Currently there is not a large economic incentive for industry to iterate at this scale.
Potential Solution: Government and academic investments to produce such facilities.
Process monitoring and control for modular and on-site (i.e., point-of-need) biomanufacturing, incorporating domain expertise.
Bottleneck/Challenge: Characterization of performance metrics for materials within context of application.
Potential Solution: Spectroscopy methods that enable high-throughput analysis of materials characterization.
Establishment of a robust network (25+) of pilot scale facilities for the production of biomolecules and biomaterials.
Bottleneck/Challenge: Currently there are very few facilities in the US that provide fermentation and downstream processing capabilities.
Potential Solution: Regional facilities that enable the utilization of various feedstocks and increase accessibility.
Bottleneck/Challenge: Pilot scale facilities need to be self sustaining.
Potential Solution: A robust number of academic, industrial, and government led projects continuing for the next decade will allow for such facilities to gain traction.
Full integration of unit operations for homogeneous biological and heterogeneous chemical and biological catalysis with separations.
Bottleneck/Challenge: Coordinated technologies and platforms for bioprocessing.
Potential Solution: Optimization and molecular design of hydrogel materials, bioreactor platforms and integration of separations.
Potential Solution: Incorporate process monitoring and control schemes.
Development of point of need production of biomolecules and biomaterials.
Bottleneck/Challenge: Insufficient demand for such infrastructure due to the way supply chains are handled and the fact that the bioeconomy is still a small section of the overall industrial base.
Potential Solution: Development of less expensive and more modular equipment will increase accessibility of fermentation and downstream processing equipment to be stood up at the point-of-need.
Quick infrastructure stand-up at point-of-need for large scale production of biomolecules and biomaterials.
Bottleneck/Challenge: The investment to stand up a biomanufacturing facility with downstream and feedstock processing is timely and costly.
Potential Solution: Development of chemical engineering processes that are more flexible and modular for downstream processing.
Potential Solution: Development of fermentation equipment that is disposable or recyclable.
Potential Solution: High titer yields would require a much smaller scale of production.
Footnotes
- Jia Liu, J., Kim, Y.S., Richardson, C.E., Tom, A., Ramakrishnan, C., Birey, F., Katsumata, T., Chen, S., Wang, C., Wang, X., Joubert, L., Jiang, Y,, Wang, H., Fenno, L.E., Tok, J.B.H., Pașca, S.P., Shen, K., Bao, Z., & Deisseroth, K. (2020). Genetically targeted chemical assembly of functional materials in living cells, tissues, and animals. Science, 367(6484), 1372-1376. https://doi.org/10.1126/science.aay4866
- Deepankumar, K., Shon, M., Nadarajan, S.P., Shin, G., Mathew, S., Ayyadurai, N., Kim, B., Choi, S., Lee, S., & Yun, H. (2014). Enhancing thermostability and organic solvent tolerance of ω‐transaminase through global incorporation of fluorotyrosine. Advanced Synthesis & Catalysis, 356(5), 993-998. https://doi.org/10.1002/adsc.201300706
- Panganiban, B., Qiao, B., Jiang, T., DelRe, C., Obadia, M.M., Nguyen, T.D., Smith, A.A.A., Hall, A., Sit, I., Crosby, M.G., Dennis, P.B., Drockenmuller, E., Olvera de la Cruz, M., & Xu, Ting. (2018). Random heteropolymers preserve protein function in foreign environments. Science, 359(6381), 1239-1243. https://doi.org/10.1126/science.aao0335
Last updated: January 19, 2021
